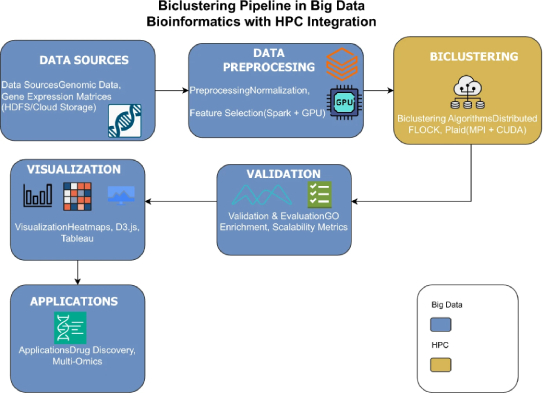
The article, titled ‘Biclustering in bioinformatics using big data and High Performance Computing applications: challenges and perspectives’, recently accepted for publication in The Journal of Supercomputing, presents a critical analysis of the state of the art in biclustering, its applications in bioinformatics, and the opportunities offered by its parallelisation on HPC platforms.
📄 Access the full article here: https://doi.org/10.1007/s11227-025-07563-6
Towards scalable and parallelised bioinformatics
Biomedical data analysis has evolved enormously with the emergence of omic technologies such as RNA-Seq, which generate massive volumes of highly dimensional data. In this context, biclustering has proven to be a particularly useful tool by allowing the detection of coherent local patterns between subsets of genes and conditions, unlike traditional clustering.
This paper provides a systematic and critical review of the most relevant biclustering techniques in bioinformatics, focusing on:
🔍 Fundamental aspects of biclustering
The theoretical foundations, their applications to real-world problems (such as biomarker identification), and their main advantages over other exploratory data analysis methods are discussed.
⚙️ Current computational challenges
The authors detail the challenges of applying biclustering to large datasets, such as computational complexity, the presence of noise, biological variability, and the need for robust validation of results.
🚀 Application in HPC and Big Data environments
One of the central themes of the paper is the need to adapt these methods to high-performance computing paradigms, such as GPU-based architectures, distributed systems with Spark or Hadoop, and parallel environments with MPI/OpenMP.
📊 Frameworks, tools and taxonomy
An updated taxonomy of existing algorithms and tools is presented, including a comparative analysis of their approaches and limitations, and a guide is proposed for their selection and adaptation depending on the type of data and resources available.
🔮 Future prospects
The article concludes with a vision for the future that highlights the need for the evaluation and benchmarking standards, adaptive and scalable algorithms, integration with AI techniques, and interoperable platforms for reproducible research.
This contribution represents a significant milestone for the scientific community working at the frontier between bioinformatics, data science and advanced computing, and lays the foundations for the development of more efficient, reproducible and scalable solutions for the analysis of complex biomedical data.
Authors:
- Aurelio López-Fernández (Pablo de Olavide University).
- Fracisco A. Gómez-Vela (Pablo de Olavide University).
- Domingo S. Rodríguez-Baena (Pablo de Olavide University).
- Fernando M. Delgado-Chaves (University of Hamburg).
- Jorge Gonzalez-Domínguez (Universidade da Coruña, Spain).




